Peripheral blood plasma collection
Plasma was collected using standard clinical procedures by drawing blood into purple-top K2-EDTA anti-coagulation vacutainer tubes. Samples were centrifuged at 1400g for 20 min without brake to separate plasma from other blood compartments. Plasma was then transferred to Cryovials for storage at -80 C. For plasma collection, samples were processed, aliquoted and frozen within 4 hours of blood draw.
Considerations for Sample Batches
For submission of samples to Olink for analysis, we thawed previously frozen plasma samples, and aliquoted samples into 96-well plates according to Olink's recommended protocols. To reduce the effect of plate-to-plate variation that could affect analysis, we attempted to keep longitudinal samples from the same subject on the same 96-well plate in our aliquot designs. We also sought to balance multiple-plate submissions by attempting to evenly distribute subjects by cohort, biological sex, and ethnicity across plates, and to reduce positional effects that could confound downstream analyses by randomizing the position of samples within each plate.
Each plate also contained some or all of a set of independent batch control samples. These samples were drawn from Healthy, Rheumatoid Arthritis, Multiple Myeloma, and Melanoma subjects in order to represent some of the biological heterogeneity in our study samples. In each batch, aliquots distributed from the same set of these samples were included to enable cross-batch bridging normalization, as described below.
Olink experiment overview
Plasma proteomic measurements were obtained from the Olink Explore 1536 and 3072 tests, comprising proteins involved in four major biological domains: cardiometabolic function, inflammation, oncology, and neurology. The Olink Explore platform (Wik, et al., 2021) utilizes “proximity extension assay” (PEA) technology, wherein target proteins are bound in solution by pairs of antibodies with complementary oligonucleotide sequences. More details about the Olink assay itself can be found on the Olink website. Hybridized oligonucleotides are then amplified and measured by next-generation sequencing, and are compared to internal, Olink-provided control samples to produce the relative quantitation termed “normalized protein expression” (NPX). Reported NPX values are log2 transformed count values that have undergone a series of normalization steps based on internal controls and average plate intensity to allow for direct comparison within a dataset. Data spanning multiple batches must be normalized using independent samples, as described below.
Batch normalization with bridging controls
A panel of independent control samples - usually 12 samples in total from Healthy, Rheumatoid Arthritis, Multiple Myeloma, and Melanoma subjects - were included in each batch, and were used to bridge all batches to a single reference batch using Olink’s recommended procedure. Briefly, for each assay and each bridge control, the difference in NPX between the query batch and the reference was calculated. The median difference for each assay (i) across all bridge controls (j) was taken as the assay batch offset:
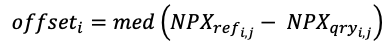
To compute the bridged NPX values, these assay batch offsets were applied to all samples in the query batch to obtain values comparable to the reference:
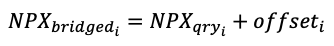
Results with warning flags were excluded from the batch offset calculation.
References
Wik L, Nordberg N, Broberg J, Björkesten J, Assarsson E, Henriksson S, et al. Proximity Extension Assay in combination with next-generation sequencing for high-throughput proteome-wide analysis. Mol Cell Proteomics. 2021;20: 100168.
doi:10.1016/j.mcpro.2021.100168