As this was our first use of the 10x Flex v2 96 sample barcode kit, we carefully assessed the quality of data aligned by Cell Ranger prior to data analysis. Cell Ranger Web Summary reports for these wells can be downloaded, and additional plots and summary statistics are available below.
IL-6 + JAK/STAT inhibitor Flex v2 Web Summaries
| File Name | Description | Download Link |
|---|---|---|
| il6_jak-stat_well_1_web_summary.html | Cell Ranger QC Summary for Well 1 | |
| il6_jak-stat_well_2_web_summary.html | Cell Ranger QC Summary for Well 2 | |
| il6_jak-stat_well_3_web_summary.html | Cell Ranger QC Summary for Well 3 | |
| il6_jak-stat_well_4_web_summary.html | Cell Ranger QC Summary for Well 4 |
Alignment and count QC
Alignment of our 10x Genomics Flex v2 libraries was performed using a pre-release beta version of Cell Ranger multi v9.1.0 to enable use of 96 sample barcodes. This functionality is now incorporated into the publically available version of Cell Ranger.
After alignment, we examined barcode alignment, count, quality, and cell barcode statistics.
Read counts, cell counts, and reads per cell:
| GEM-X Well ID | Reads | Cells | Reads per Cell |
|---|---|---|---|
| PAC1W1 | 5,372,953,653 | 1,050,355 | 5,115 |
| PAC1W2 | 5,801,847,691 | 1,094,469 | 5,301 |
| PAC1W3 | 5,822,737,681 | 1,123,811 | 5,181 |
| PAC1W4 | 4,689,840,447 | 969,171 | 4,839 |
Read saturation, mapping, and quality:
| GEM-X Well ID | Saturation | Percent Mapped | Percent Passing QC |
|---|---|---|---|
| PAC1W1 | 72.5 | 97.6 | 96.7 |
| PAC1W2 | 72.7 | 97.5 | 96.6 |
| PAC1W3 | 72.7 | 97.6 | 96.7 |
| PAC1W4 | 70.6 | 97.7 | 96.7 |
UMI and Gene QC across wells and replicates
Our experiment utilized 4 wells on a GEM-X capture chip, each loaded with the same pool of cells. This enables us to assess consistency of QC metrics between wells.
We also had replicate drug treatment wells within our 96-well experiment plates: 4 wells were unstimulated controls, treated with neither IL-6 nor a JAK/STAT inhibitor. 4 additional wells were treated only with IL-6 stimulation, without JAK/STAT inhibitors.
To assess quality, we first looked to the UMI and Gene Detection distributions across wells. Frequent users of whole-transcriptome scRNA-seq approaches may notice low UMI and Gene detection values relative to standard 10x assays - this is due to the use of a limited panel of 1,916 gene probes used in this pre-release assay.
Across all 4 wells, we found consistent UMI and gene detection:
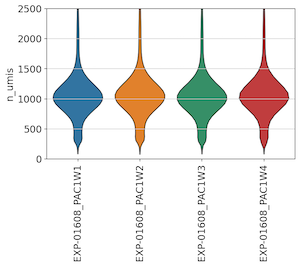
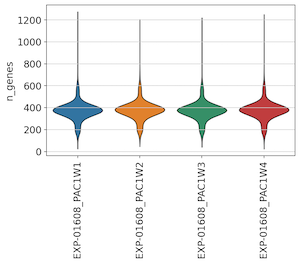
Across Unstimulated and IL-6 control replicates, we also observed consistent UMI and gene detection:
UMIs per cell across replicate wells:
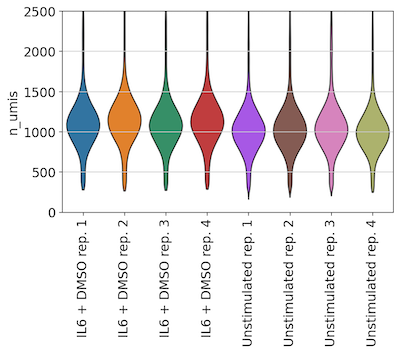
Genes per cell across replicate wells:
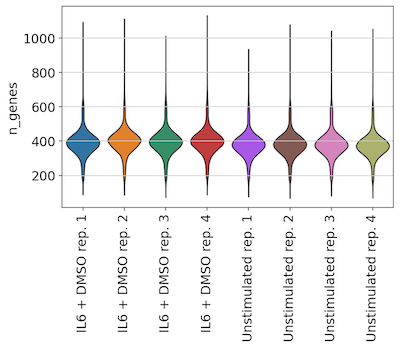
Effects of overloading
The 10x Genomics GEM-X Flex v2 User Guide (CG000834, Rev A) provides cell loading guidelines for use of up to ~1.4 million cells per well with 96-sample multiplexing. We were able to load ~1.2 million cells, and recovered ~1 million cells per well.
This involved relying on substantial GEM "overloading" in which many cells are encapsulated in the same GEM, and the gene barcodes from these cells are deconvoluted using the 96 sample barcodes. Because of the high number of sample barcodes, even when cells are overloaded, there is a low probability that cells from the same sample will be captured in the same GEM, resulting in a non-deconvoluted doublet.
To assess the actual extent of overloading in our assay, we used the cell barcodes from Cell Ranger outputs to assign the original GEM barcode to each cell in each well of our experiment, and checked the distribution of cells per GEM:
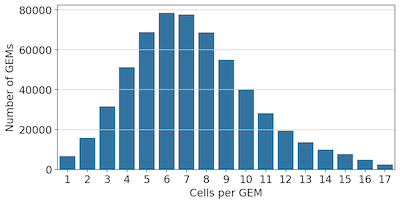
We can see that the vast majority of cells in our experiment were in an overloaded GEM. To see how this effects data quality, we grouped cells by the number of Cells per GEM, and checked the distribution of UMIs and genes detected:
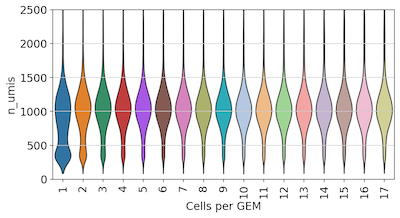
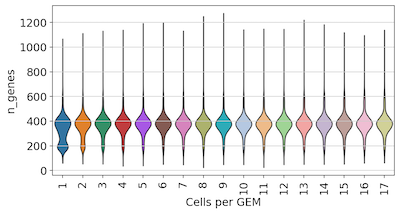
Surprisingly, we found no strong effect on UMI or gene detection based on the number of cells per GEM. We do observe some cells with low UMI or gene detection in GEMs with only 1-2 cells. These may be low-quality or dying cells encapsulated individually or with only one other cell.