Formalin fixed parafin embedded (FFPE) human tissue samples from the duodenum region of small intestine and from skin were provided by the lab of Prof. Laura Mackay at the Doherty Institute.
Atera profiling of tissue sections was carried out by 10x Genomics, including sample perparation, probe hybridization, ligation and amplification, staining, and fluorescent probe hybridization, imaging, and decoding:
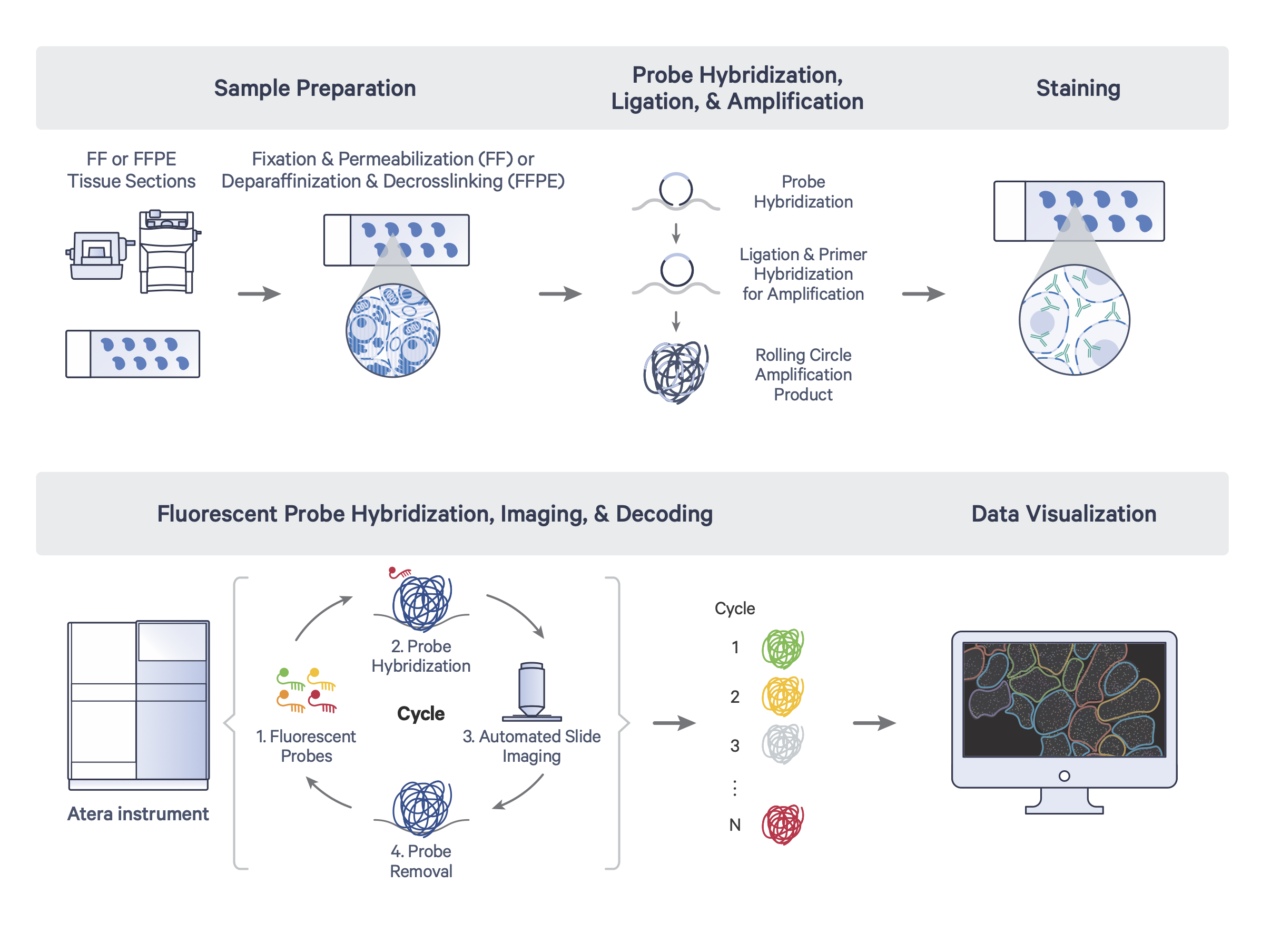
Atera workflow diagram courtesy of 10x Genomics
After receiving data from 10x Genomics, we assessed basic quality control metrics for each tissue, utilizing the cell_feature_matrix.h5 and cells.csv.gz outputs provided by 10x Genomics.
Human duodenum quality and labeling
Based on segmentation provided by 10x genomics, our human duodenum sample provided data from 320,885 cells. We observed 3,291 mean transcript counts per cell (st. dev = 2,716; median = 2,641). Gene detection was similarly high: 1,745 genes per cell on average (st. dev = 1,078; median = 1,603).
For labeling, we filtered cells with < 150 transcript counts (9,405 cells) and retained 311,480 cells for downstream steps (97% retained).
To apply cell type labels, we utilized the scanpy functions scanpy.pp.normalize_total() with the parameter target_sum = 1e4, followed by scanpy.pp.log1p(). We performed label assignment using the CellTypist package, and utilized the cell type model for Intestine provided as part of the CellHint auto-harmonised and auto-integrated Organ Atlas resource (specifically, the "All combined model"). Cell type prediction was performed using the celltypist.annotate() function with the majority_voting = True parameter. Majority voting labels were used as a first pass for label assignment, followed by manual curation to combine highly related categories to identify 20 initial cell types. We expect further analysis will yield confident, high-resolution cell type assignments.
To visualize cell type labels in a 2-dimensional projection, we further processed the data using scanpy to identifiy 2,6222 highly variable genes with scanpy.pp.highly_variable_genes(), followed by dimensionality reduction with scanpy.tl.pca(), nearest neighbor analysis with scanpy.pp.neighbors() with the parameter n_pcs = 30, and UMAP projection with scanpy.tl.umap(). We were then able to plot cell type assignments on UMAP coordinates:
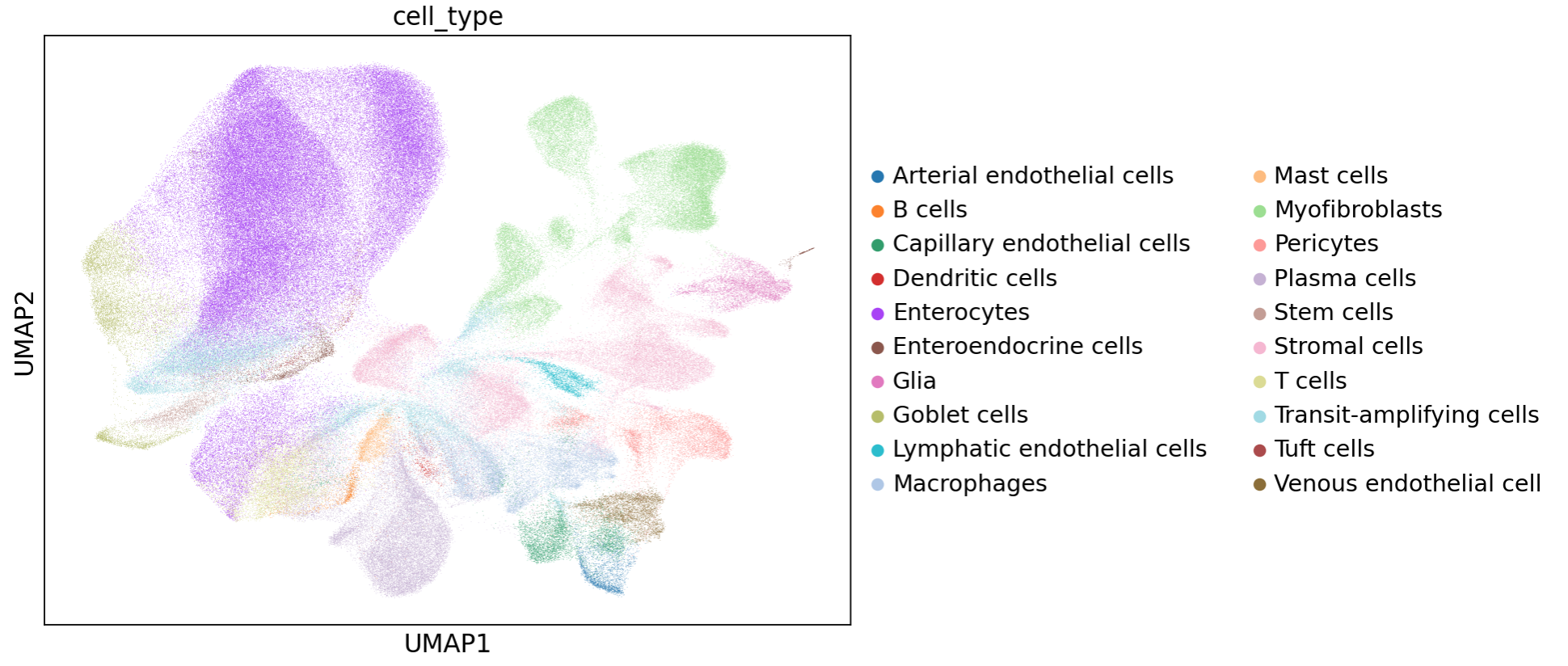
After projection, we were able to identify distinct markers for most of these duodenum cell types thanks to the large transcriptomic coverage provided by the Atera data:
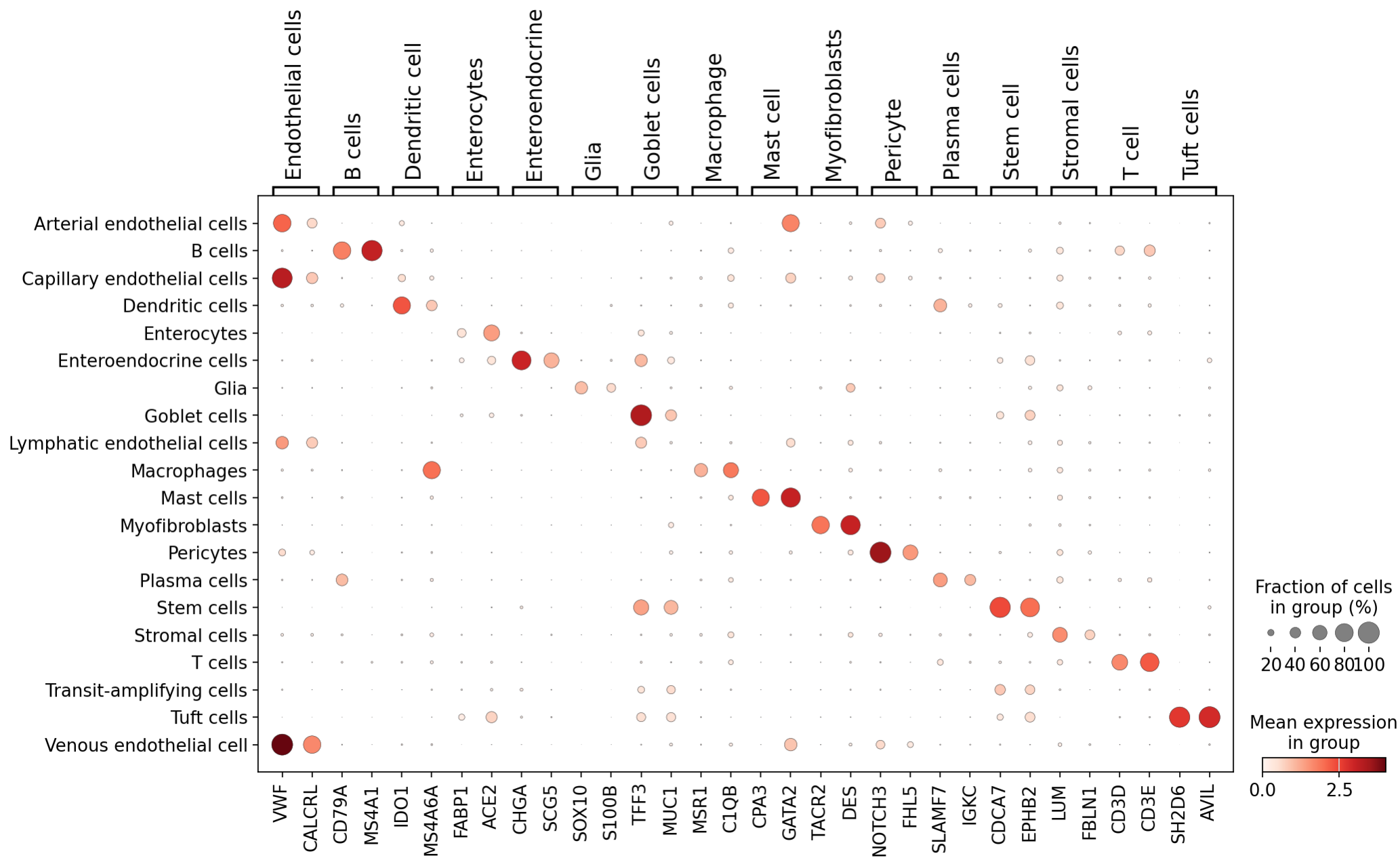
Additional inspection of cell types was carried out by visualization of cell locations in situ - see the interactive visualization tools on the main page of this report.
After labeling, we inspected the distributions of transcript and gene detection across duodenum cell types.
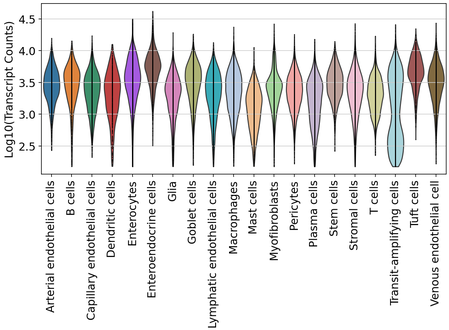
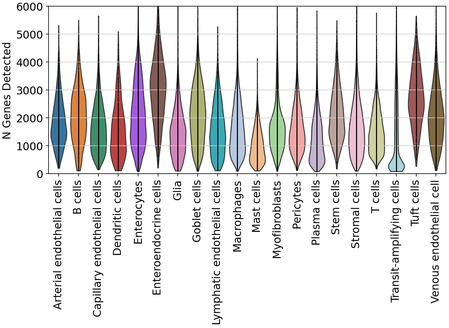
Human skin quality and labeling
As for duodenum, we carefully assessed the quality of our data generated using a human skin sample. As expected for this tissue type, we observed many fewer segmented cells, as the cell-rich epidermis layers made up a minority of the total area of this tissue sample.
Segmentation provided by 10x Genomics allowed analysis of 42,851 cells, with mean detection of 3,757 transcripts (st. dev. = 2,817; median = 3,121) from 1,934 genes (st. dev. = 1,049; median = 1,847). After filtering for cells with > 150 transcripts, we removed 1,309 cells and retained 41,542 for cell type labeling (97% retained).
As described for duodenum, labeling was performed using CellTypist predictions with majority_voting = True. For skin, we utilized the Adult_Human_Skin model provided on the CellTypist website in the Models section. After prediction, majority voting, and label merging, we identified 15 cell types. As for duodenum, we performed dimensionality reduction to visualize these cells:
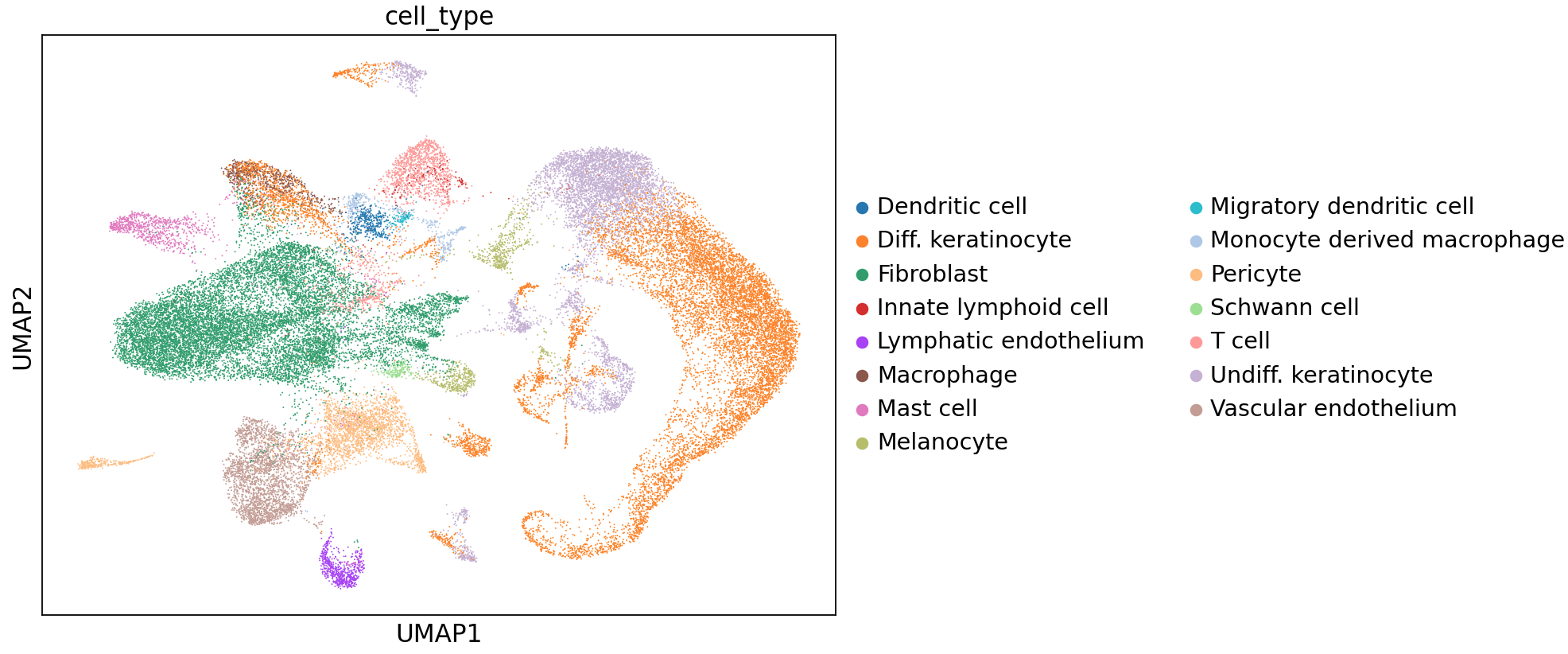
We were able to find marker genes to distinguish these cell types in the whole-transcriptome dataset:
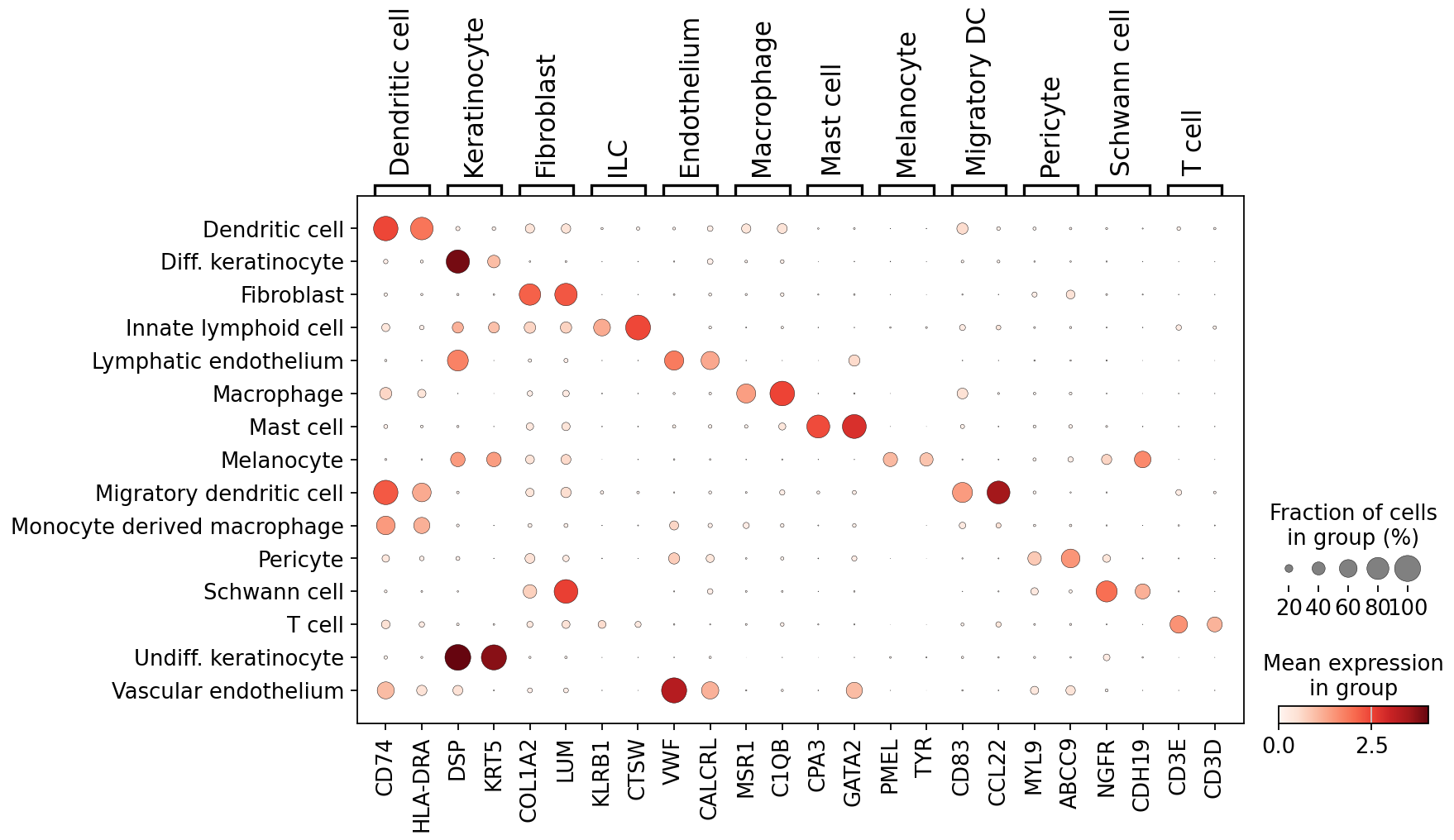
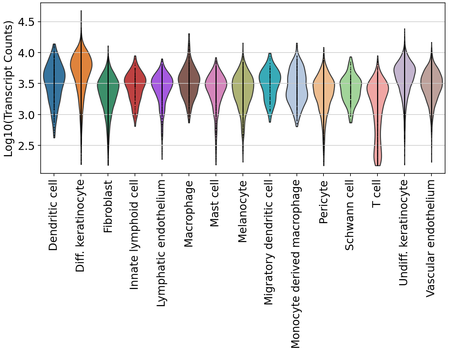
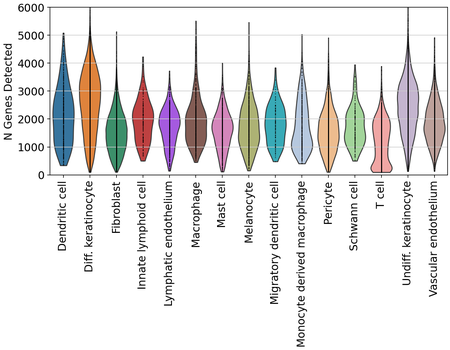
Finally, we also examined transcript and gene detection across skin cell types. We see variation in transcript and gene detection across cell types, which will need to be further examined.