Use the Get File Descriptors SDK Method (Tutorial)
Last updated 2026-05-07At a Glance
This document explains how to use get_file_descriptors()(Python) | getFileDescriptors (R) to retrieve file-level metadata based on query criteria. It accepts a query dictionary with list-based filters and returns a dictionary of DataFrames, including descriptors, lab results, specimens, and survey data. You can query either public or private files using the is_public flag, but you must include fileType in the query when accessing nonpublic data. If you have questions or need help, contact Support.
When to Use This Method
Use get_file_descriptors() | getFileDescriptors() to load all file descriptors (file/sample/subject metadata) for a given file type:
Retrieve structured metadata about files that match specific criteria
Explore associated data like lab results, specimens, or survey responses alongside file descriptors
Build workflows that depend on filtering files by attribute (for example, file type or project)
Analyze or export file metadata in a tabular format for downstream processing
Establish a consistent retrieval method to work with public datasets or authenticated project data
SDK Method
The get_file_descriptors() signature is shown in the box below, and the method parameters follow. Click the tabs to toggle from Python to R.
Signature
Python signature
get_file_descriptors
Retrieves file descriptors based on user's query.
hp.get_file_descriptors(
query_dict: dict = None,
is_public: bool = False
)R signature
getFileDescriptors
Loads all file descriptors (file/sample/subject metadata) for a given file type.
getFileDescriptors(
fileType,
filter = NULL,
toDF = FALSE
)Parameters
Python parameters
Parameter | Data type | Required or optional | Description | |||
|---|---|---|---|---|---|---|
|
| required | Dictionary object containing search parameters using Mongo query language. Default is | |||
|
| required | If |
R parameters
|
|
| Description | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
|
| optional | Type of file to search for (e.g., "scRNA-seq-labeled", "FlowCytometry-supervised-stats"). | |||||||
|
| required | List filter to narrow the search. Same format as | |||||||
|
| required | Logical operator indicating whether to return the results as a list of |
 Get Help
Get Help
If you get stuck during a get_file_descriptors() | readSubjects call, refer to the steps of this tutorial (examples are in Python unless otherwise specified). To use the baked-in help in your IDE, try one of the following commands. Still not working? Contact Support.
Python | R | Output | ||
|---|---|---|---|---|
|
| Function signature, list of parameters, class, and a brief description of the method in a compact plain-text format | ||
|
| Method signature, docstring (description), file location, and file type in more readable format | ||
|
| Signature, docstring, file path, a verbose set of metadata, and the source code for the method |
Instructions
Define a query
Navigate to HISE, sign in , open an IDE, and set up your environment. For details, see Create Your First HISE IDE (Tutorial) and Use HISE SDK Methods and Get Help in the IDE .

Use a query dictionary to specify which files to describe. All filter values must be lists. This example filters the returned file descriptors by file type, panel, and cohort GUID.
In Python, you can include multiplefileTypevalues in the list to retrieve several file types in a single call. The RgetFileDescriptors()method accepts only onefileTypeper call.
query_dict = {
"fileType": ["FlowCytometry-labeled-expr-csv"],
"panel": ["PT1"],
"cohortGuid": ["UP1"]
}
NOTE
If you prefer, you can pass the dictionary contents directly in your get_file_descriptors() call. The approach shown above is recommended, however, because it lets you confirm the values before passing them in, reuse the query_dict in other SDK methods, or modify a single key and rerun the call as you iterate on your query.
df_dict = hp.get_file_descriptors(
query_dict={
"fileType": ["FlowCytometry-labeled-expr-csv"],
"panel": ["PT1"],
"cohortGuid": ["UP1"]
}
)Get file descriptors
To retrieve file‑level metadata and related data frames, call
get_file_descriptors()with your query.df_dict = hp.get_file_descriptors(query_dict=query_dict) df_dict.keys()If matching files are found,
df_dict.keys()returns entries that match the tables available for your query.
"descriptors"
"labResults"
"specimens"
"survey"
2. View the file descriptors table:descriptors_df = df_dict["descriptors"] descriptors_df.head()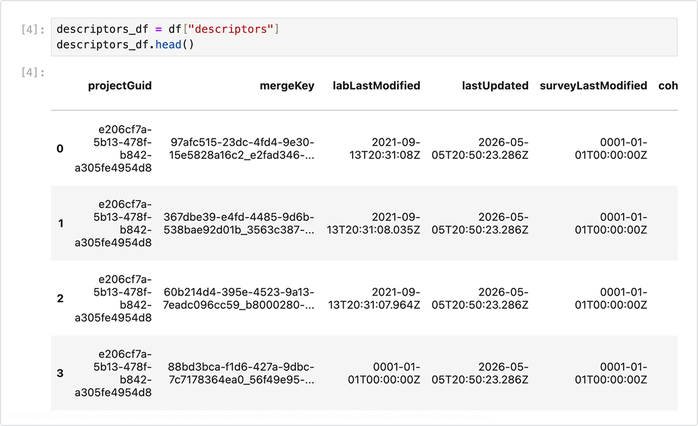
Cache matching files (optional)
To work with the matching files in your IDE (for example, to open them or pass them to another analysis function), cache them locally using their file IDs.
Retrieve the files that match your query. You can use these local paths in downstream analysis or other SDK calls.
file_ids = descriptors_df["file.id"].tolist() cached_paths = hp.reader.cache_files(file_ids) cached_paths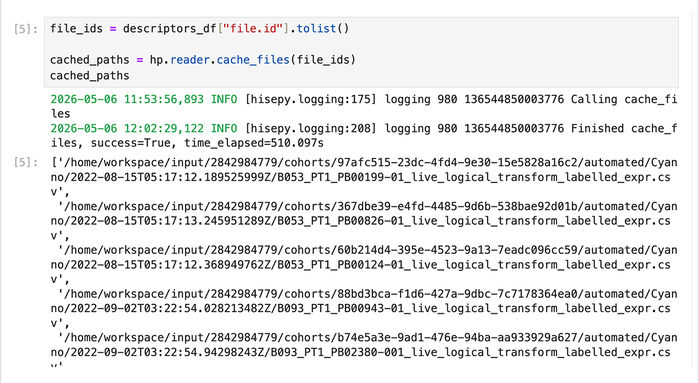
 Related Resources
Related Resources
Query SDK File Type (Tutorial)
Use the Read Files SDK Method (Tutorial)